5 Minimalist Tips for Data Scientists to reduce frustration while working with Pandas
2022-02-06
[Edit]
Join our Discord community, “Code Quality for Data Science (CQ4DS)”, to learn more about the topic: https://discord.gg/8uUZNMCad2. All DSes are welcome regardless of skill level!
[Edit]
I just came out of a two-week project where we needed to create a prototype in no time with unknown data. Speed was of essence, so I needed to write tons of ad-hoc analysis with notebooks and pandas and needed to brush off my knowledge. Now that I was at it, I might as well write it up for my readers.
Pandas is one of those techs that people, including me, like to complain about yet still keep using it. Pandas have the highest number of StackOverflow questions, which can be a testament to its popularity but also to its complexity.
I have used pandas for about eight years and built a set of tried-and-tested habits to keep my sanity.
For production-grade programming, I do not recommend using pandas at all. You can read more about how to maintain data science code in production here: Refactoring for Data Scientists
1.) Saving and Loading
Pandas provides a massive amount of utilities to load data; CSV, pickle, SQL, etc., you name it. I think this is one of the reasons for its success: it is easy to start working with it.
On the other hand, to simplify this, it usually “silently” makes assumptions about your data and converts it to whatever type it thinks you wanted (often incorrectly). If this is a problem (or just doing it anyway), I use pydantic based data modelling and loading:
It provides a lot of saving functions as well, though I would make a suggestion here. If you need to save pandas dataframes, you are probably doing some work that should happen outside of notebooks in a pipeline with a well-defined data model.
Anyway, if I need to, I use pickling to make sure that if, for whatever reason, I save something, I get back exactly the same object as I saved.
(Don’t be surprised about the short/single-letter variable names in the examples. I use them; you use them, everyone uses them. What’s the point of fighting this habit? It’s fine in throwaway/analytics code but do refactor them if you move your code to production.)
2.) Speed vs readability
If you are working in a notebook and with pandas, you are usually limited by the memory of your own machine (often your own laptop). Performance is rarely a relevant concern for you, and your efforts should be focused on readability and changeability as ad-hoc analysis demands.
As we all know: “Premature optimisation is the root of all evil”.
Despite this, we are all impatient and hate waiting and want to move faster. This is a natural desire, so to counter that, I start all cells with the %%time magic and use tqdm on loops and long groupby/apply situations:
Apparently, this needs tqdm=4.62.2, so update your packages.
You will see the pointlessness of optimisation if you see sub 1 minute waiting times, and you realise that you would have sacrificed readability for that.
3.) Creating a new column
I always create columns one by one, so their generation is not coupled, and I can change them individually. For these, I have three small patterns.
Single-column and zipping are pretty efficient, though not very scalable if you want to work with many columns. Itertuples has a good baseline performance. It also has the additional advantage that the code you write in the processing function will treat `row` as an object-like entity. This will help you refactor it if you need to move the code to prod. I try to use less zipping and more itertuples for that reason.
If you still feel it’s slow, just use tqdm as the example shows.
4.) Groupby and generate multiple columns
If you do any kind of descriptive analytics, you need to do groupbys all the time, so you would expect this is a well-designed part of the API, and you would have learnt it well by now.
Despite this, every single time I needed to do a groupby, I had to go to StackOverflow, breaking my flow and thought process. Very annoying. I write some code and run it, but the output is not what I expected.
There are a thousand ways to do this based on the input, the columns, the apply method, and the requested outputs. I have no time for this while my head is thinking about some domain/data problem. I am not going to fight my tools.
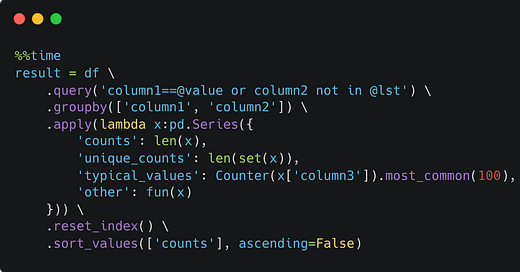
So I decided to simplify this to a single way to do it. And no matter how simple the problem is, I always do it this way. It massively reduces cognitive load so I can focus on the task:
This ensures that the result will have all the columns from the grouping column set and the ones in the lambda expression. The fun() function will be passed a small dataframe which I usually process with itertuples and return a list.
Anything more complicated, and you are probably better off with productionised code with well-defined input/output data types (see above at Point 1).
5.) Merging and joining
These bit me so many times that I pretty much stopped using them. I either need to dig documentation/SO for answers, or I overlook some parameter, and pandas silently creates an incorrect output.
If I need to do something involving more than one dataframes, I’d instead write it out as python code and use some of the patterns from above:
Create a result set with groupby from Point 4.
Extract the relevant data by some key into a lookup table (plain old dictionary).
Create a new column in the original dataframe with Point 3.
If this fails … StackOverflow …
+1.) Quick check with Counter
I often need to quickly check something in a column as part of EDA. I use Counter very heavily (and you can see one typical use in Point 4 at the groupby example):
Summary
I try to simplify the number of tools I use when analysing. I often don’t need to worry about performance because if I do, I would usually not use notebooks. If I am still bothered by slowness, I use the %%time magic and tqdm.
The few available tools enable me to move beyond the task of writing code. I can think of abstract analysis problems, and there is almost always just one way to solve them so I can type automatically, reducing cognitive load.
If you found this list helpful, please share it with others to learn about it. Subscribe for more content like this in the future:








Hi Laszlo, for Point 5, why not using df['column'].value_counts(dropna=False).head(n=100)? Won't it give you the same thing along with a distribution?